Lions, Tigers, and Bears, Oh … and Hominins?
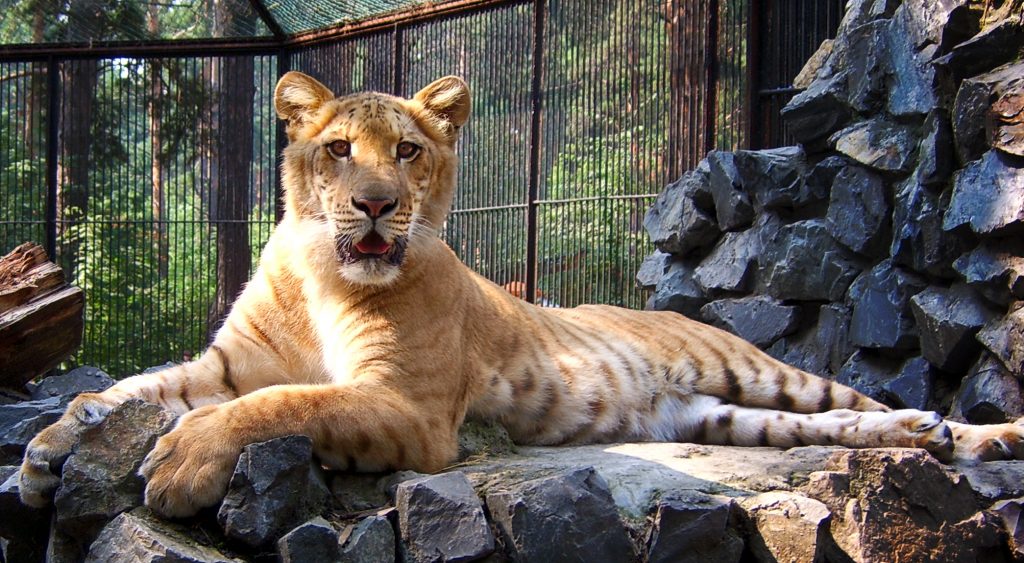
Ligers, tigons, and pizzlies, oh my! What do these critters have in common? They are the offspring of animals that belong to distinct lineages, or populations that have diverged evolutionarily. In other words, they are hybrids. A liger is produced when a male lion and a female tiger mate. A tigon is the offspring of a male tiger and a female lion, and a pizzly, as cute as it sounds, is a hybrid of a grizzly bear and a polar bear—so it isn’t an animal you’d want to cuddle.
Hybridization is fascinating from multiple perspectives. It provides insight into reproduction, genetics, and behavior. Researchers who study hybrid animals can also examine the relationships among habitat distribution, ecological niche, and population structure by tracking what goes on in “hybrid zones” where populations meet, individuals interbreed, and genes flow. Richard Harrison, a Cornell University evolutionary biologist and pioneer in his field, who recently passed away, described hybrid zones as “natural laboratories of the evolutionary process.”
Hybrids are often a key element in discussions about species concepts. Biologist Richard Mayden identified 24 species concepts, and John S. Wilkins, a historian and philosopher of science, provided 26 definitions of species.
Species concepts matter because the application of a particular species concept determines how a species is characterized through space and time. For example, as Emily Graslie nicely explains in The Brain Scoop, the biological species concept would put polar bears and grizzly (brown) bears in the same species because they can mate and produce fertile offspring (Wilkins’ “biospecies”); however, the ecological species concept would designate them as separate species due to differences in their habitat-related adaptations (Wilkins’ “ecospecies”). The phylogenetic species concept would also assign the bears to separate species (Wilkins’ “phylogenetic taxon species”) due to their distinct evolutionary histories dating back to when polar bears diverged from grizzly bears at least 150,000 years ago. In 2012, a genetic analysis of a combined 35 populations of polar bears and grizzlies confirmed there has been little to no gene flow (hybridization) between them in the wild and that differences in their DNA were consistent with the two bears having separate gene pools and therefore being different species—the genetic species concept (Wilkins’ “genetic species”).
And the application of species concepts has implications for how researchers interpret hominin species, including those in the genus Homo—even Homo sapiens. As the physical anthropologist Clifford “Cliff” Jolly noted in 2001, the primary question pertaining to Homo species is often couched as either: “[Should] Neandertals be classified within Homo sapiens?” or “How many species of the genus Homo existed contemporaneously during the later Pleistocene (circa 150,000 to 30,000 years ago)?” The answer depends on the species concept and on the evidence examined.
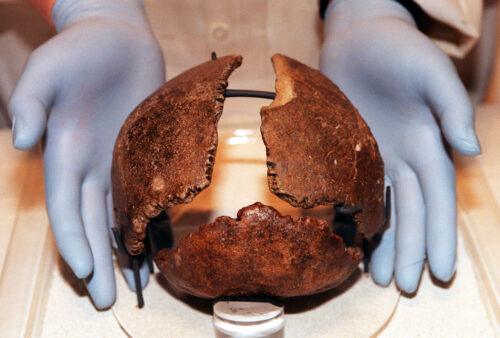
For decades, researchers addressing the “Neanderthal problem” and the matter of contemporaneous Homo species could only rely on the morphology (bones and teeth) of Homo specimens and on observations of living hybrid organisms to make inferences about interbreeding between extinct hominins. But due to advances in human genomics and ancient DNA analysis, human evolutionary anthropologists have new tools to explore and debate this and other questions about human origins.
“The ‘Neandertal problem’ has had different connotations for every generation of human evolutionists.” —Cliff Jolly
It should be noted that the “Neanderthal problem” would now more appropriately be called the “Neanderthal and Denisovan and probably-other-undiscovered-hominins problem.”
At the 85th annual meeting of the American Association of Physical Anthropologists (AAPA) in April 2016, the topic of one of the opening sessions was how the study of human evolution could be informed by an understanding of hybridization in other animals. Biologists and anthropologists shared what is known about hybridization in birds, monkeys, bears, mice, amphibians, and wild and domestic dogs.
One of the co-organizers of the session, Rebecca Ackermann, opened the symposium with an overview of questions and research relevant to hybridization in human evolution. Ackermann explained how genetic and fossil data paint a “complex picture of lineage divergence and remerger, resulting in the diverse phenotypes and genotypes we see among living people.” She set the stage for the other presentations by explaining that hybridization studies of living organisms provide an opportunity to take a fresh look at the drivers of human evolution.
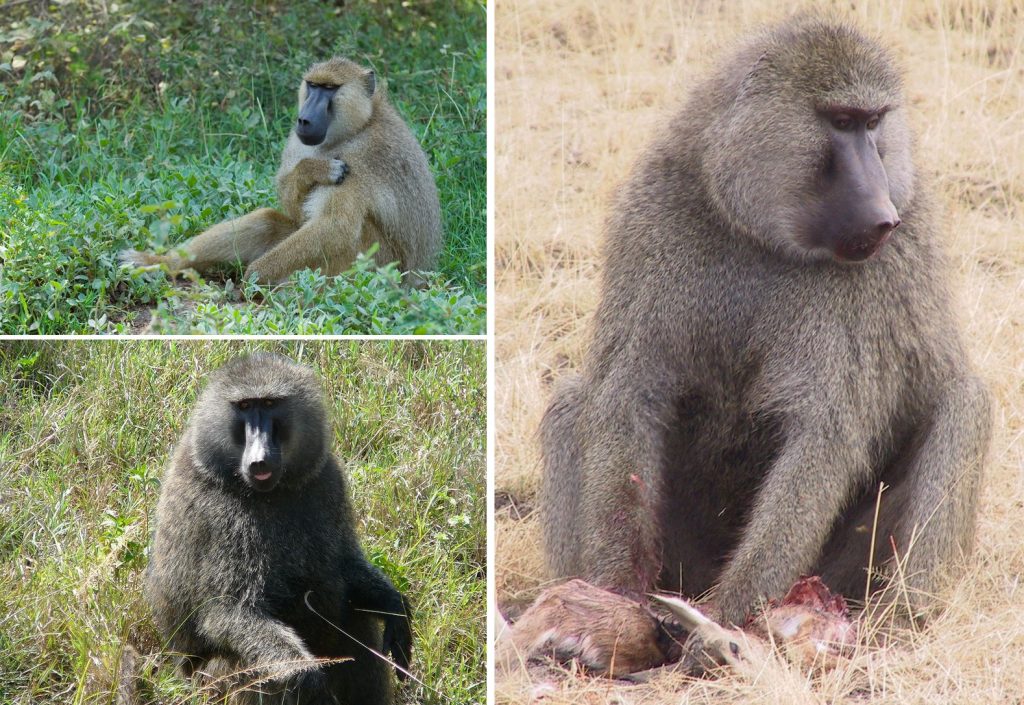
Two of the AAPA talks focused on baboons. As Jolly laid out in his 2001 publication, baboons can be viewed as living analogues for some aspects of human evolution. Some baboons shared habitats with hominins, and climatic and ecological changes may have affected the animals similarly. And gene flow between baboon populations could be a model for gene flow between hominin groups that overlapped in space and time.
One group of AAPA researchers reported on genetic analyses demonstrating that hybridization may have shaped the historical geographic distribution of baboon populations and the process of baboon speciation. Similarly, genetic studies of modern humans and of ancient DNA from Denisovan and Neanderthal specimens confirm some degree of interbreeding between these populations and contribute to the story of recent hominin evolution. But just because you know you are 2.1 percent Neanderthal doesn’t mean researchers have solved the “Neanderthal problem”—because the answer still depends on how you define a species.
Evidence from baboon hybrids also shows that hybridization may be responsible for some of the variation we see in the Pleistocene fossil record. In the final AAPA presentation of the morning, the researcher noted that the likelihood of finding a fossilized hybrid offspring of two distinct hominin populations is small; however, evidence was presented that the size and shape of at least one hominin specimen—a potential human and Neanderthal hybrid—if not others, can be explained based on the results of experiments with mouse hybrids (as well as DNA analysis). This indicates that some traits researchers use to define species in the hominin fossil record may be the result of gene flow between neighboring populations. If that is the case, just how many hominin species did live during the later Pleistocene?
Though I realize some of my readers are still contemplating whether humans are wild, I leave you with one more question, gratias ago Ernst Mayr: “Are species realities of nature or are they simply theoretical constructs of the human mind?”